2023-HGU-ML Lecture 9. Nonlinear Dimension Reduction
Nonlinear Dimension Reduction
- Kernel machine and manifold learning
- extension of linear models with the kernel trick
- kernel PCA
- kernel Fisher discriminant (kernel FD)
- Kernel trick
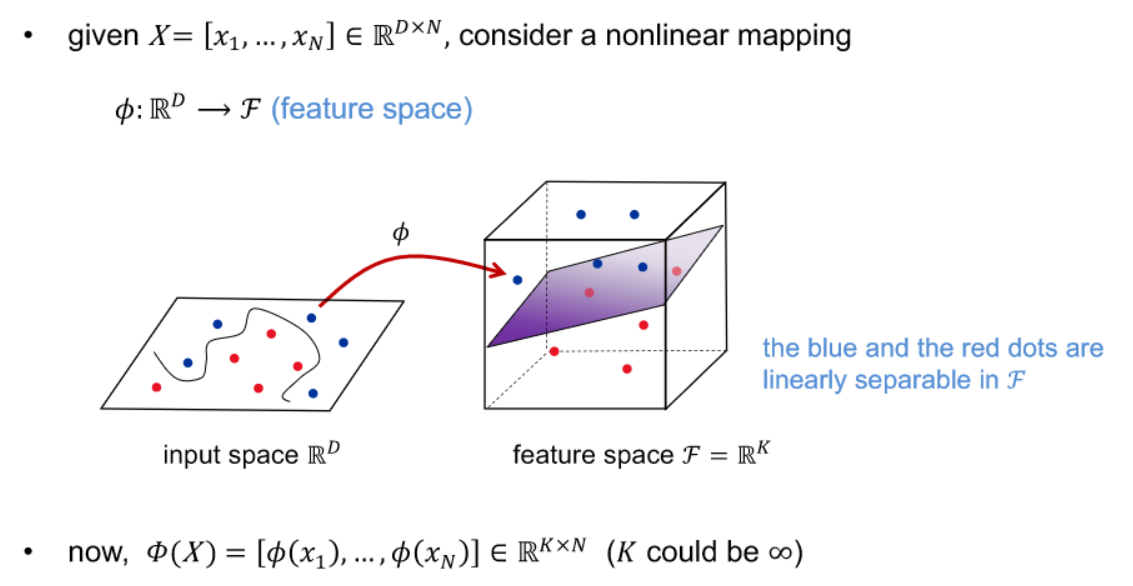
- many ML algorithms are based on relations between samples’ inner product
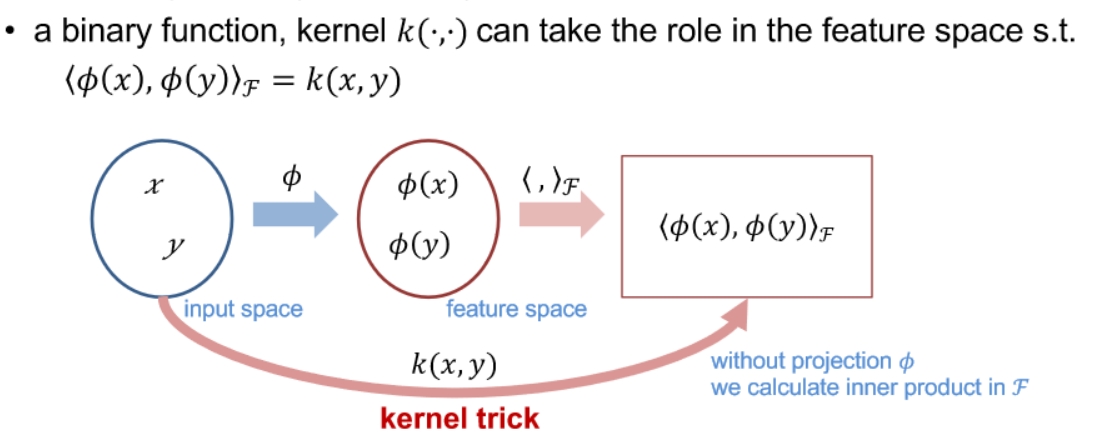
- The kernel trick is a method in machine learning and support vector machines (SVMs) that allows the application of a linear algorithm in a high-dimensional feature space without explicitly calculating the transformed feature vectors. It is achieved by using a kernel function to compute the dot product between the transformed data points, which implicitly represents the higher-dimensional space. This technique is especially powerful when dealing with non-linearly separable data, enabling linear algorithms to effectively capture complex relationships by operating in a higher-dimensional space.
- Mercer’s theorem
- any PSD kernel can be expressed as an inner product in some space
- if a kernel function k(x, y) is positive semi definite(PSD), a PSD matrix can be eigen-decomposed
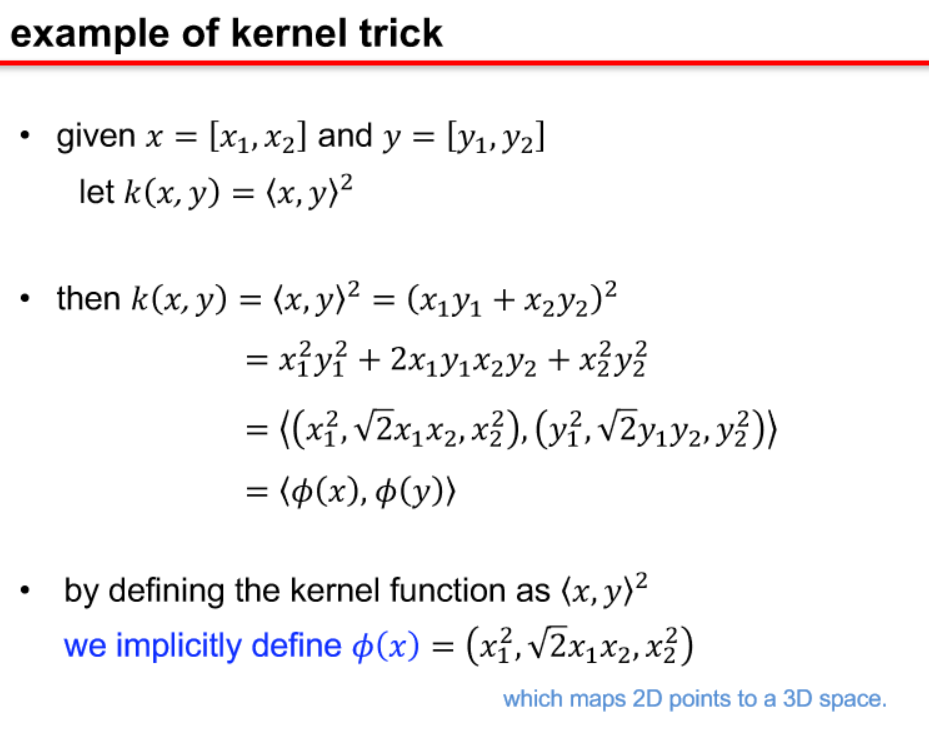

-
kernel functions
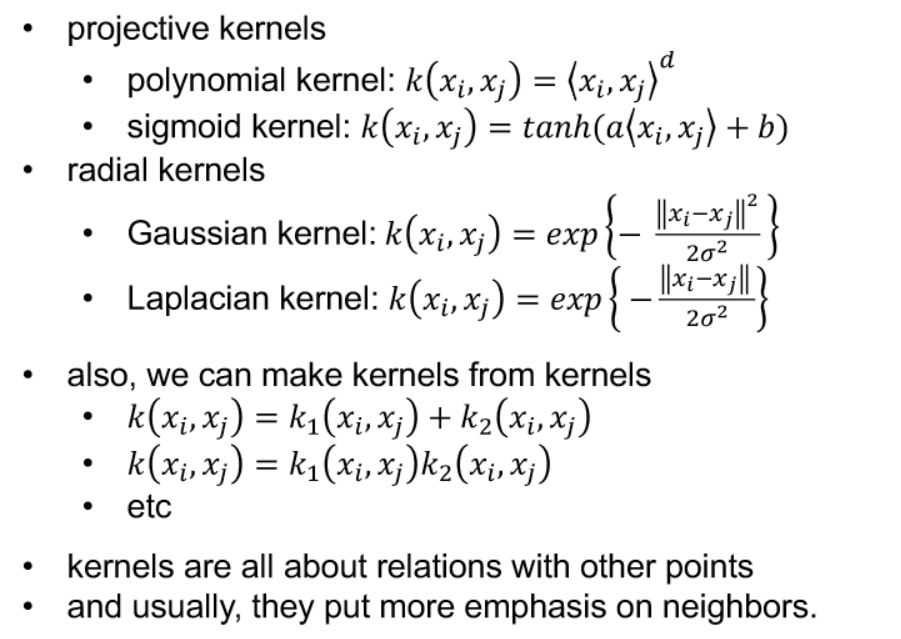

- kernel PCA
- Kernel Principal Component Analysis (Kernel PCA) is an extension of Principal Component Analysis (PCA) that utilizes the kernel trick to implicitly map input data into a higher-dimensional feature space. In traditional PCA, linear transformations are applied to find the principal components, but in Kernel PCA, a kernel function is used to capture non-linear relationships in the data. This allows Kernel PCA to uncover complex patterns and structures in high-dimensional spaces, making it particularly useful for tasks such as dimensionality reduction and non-linear feature extraction in machine learning.
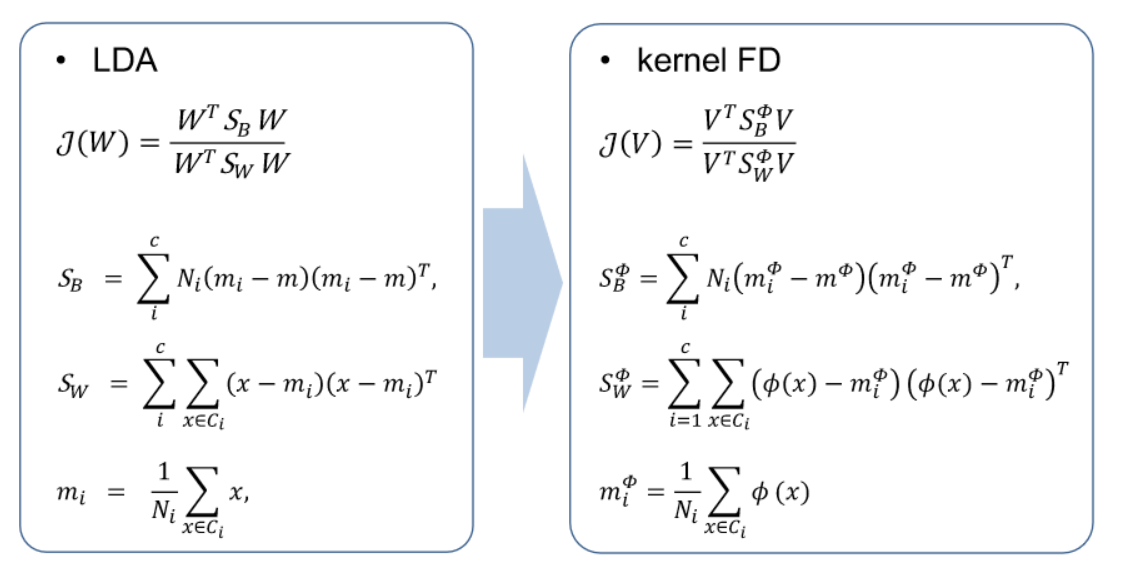
- kernel FD
- Kernel Fisher Discriminant (KFD) is an extension of Fisher’s Linear Discriminant Analysis (LDA) that incorporates the kernel trick to handle non-linearly separable data. Fisher’s LDA is a method used for finding linear combinations of features that best separate different classes in a dataset. KFD, through the use of a kernel function, allows this linear separation to be performed in a higher-dimensional space, enabling the discrimination of classes in a non-linear manner. Similar to Kernel PCA, Kernel Fisher Discriminant is particularly useful when dealing with complex, non-linear relationships in the data.
- manifold learning
- usually, nonlinear dimension reduction
- find the invariant property of a data set
- linear methods (PCA, LDA, ICA, NMF, etc)
- nonlinear methods (kernel PCA, kernel FD, Isomap, LLE, etc)

- Isomap
- manifold learning algorithms are to find the embedded manifold from data samples in a high dimensional space
- Isomap algorithm
- uses geodesic distances on a neighborhood graph in the framework of MDS
- https://en.wikipedia.org/wiki/Multidimensional_scaling

- Deciding neighbors
- KNN
- epsilon neighborhood
-
kernel Isomap

- positive semi-definiteness
- locally linear embedding
- keep the neighbors
- Locally Linear Embedding (LLE) is a nonlinear dimensionality reduction algorithm that seeks to preserve the local linear relationships within a dataset. The key idea behind LLE is to represent each data point as a weighted linear combination of its neighbors, thereby capturing the local geometry of the data. Here’s a brief explanation of the LLE algorithm:
- Local Reconstruction:
- For each data point in the high-dimensional space, LLE identifies its k nearest neighbors. These neighbors are the points that are most similar to the given point based on pairwise distances.
- Local Weight Optimization:
- LLE then seeks to reconstruct the data point by finding the weights (coefficients) that best represent it as a linear combination of its neighbors. The reconstruction is performed in a way that minimizes the difference between the original point and its reconstruction.
- Global Embedding:
- After obtaining the local linear representations for all data points, LLE seeks a lower-dimensional representation (embedding) for the entire dataset while preserving these local relationships. This is achieved by minimizing a cost function that enforces consistency in the local linear relationships across the entire dataset.
- Final Embedding:
- The resulting lower-dimensional representation is obtained by stacking the embeddings of all data points. This representation captures the intrinsic geometry of the data, emphasizing the local linear relationships.
- Local Reconstruction:
- Isomap and LLE
- Isomap takes a global strategy, while LLE local
- LLE analyzes local linear coefficients and reconstruction error
- no need to solve large dynamic programming problems
- related to spectral clustering
- stochastic neighbor embedding
- probabilistically decides if points are neighbors to a given point
- convert similarity into the probability
- neighbors are selected probabilistically
- use distances in a low dimensional space which define probabilities
- compute KL divergence between the two probabilities
- minimize KL with respect to yi
- Stochastic Neighbor Embedding (SNE) is a dimensionality reduction technique that aims to capture the local and global structures of high-dimensional data in a lower-dimensional space. SNE is particularly effective at preserving pairwise similarities between data points, making it well-suited for visualization and exploration of complex datasets.
- t-SNE
- a symmetrized version of SNE with t-Student instead of Gaussian
- t-SNE (t-Distributed Stochastic Neighbor Embedding) is an extension of the original SNE (Stochastic Neighbor Embedding) algorithm designed to address some of its limitations. Both SNE and t-SNE are nonlinear dimensionality reduction techniques that focus on preserving pairwise similarities between data points.
-
t-SNE and SNE
Comparison:
- Crowding Problem:
- One of the main challenges with SNE is the crowding problem, where points in high-dimensional space are too close together and end up being crowded in the lower-dimensional space. t-SNE addresses this issue by using a heavy-tailed distribution, leading to better separation of clusters.
- Preservation of Global Structure:
- While SNE can struggle with preserving global structures, t-SNE is designed to perform better in this aspect. It is often more effective in revealing the overall structure of the data.
- Computational Complexity:
- t-SNE can be computationally more expensive than SNE, especially for large datasets, due to the need to compute and optimize the heavy-tailed distribution
- https://woosikyang.github.io/first-post.html
- Crowding Problem:
- neural network approaches
- weakness in manifold learning (kernel machine): it does not scale well with sample size. (scales quadratically with the size) the training data is referenced for test data
- let’s get back to parametric models, especially neural networks
- self organizing map
- restricted Boltzmann machine
- autoencoder (trained with backprop)